Breadcrumb
- Home
- Research
Research
Cryptococcal Dissemination
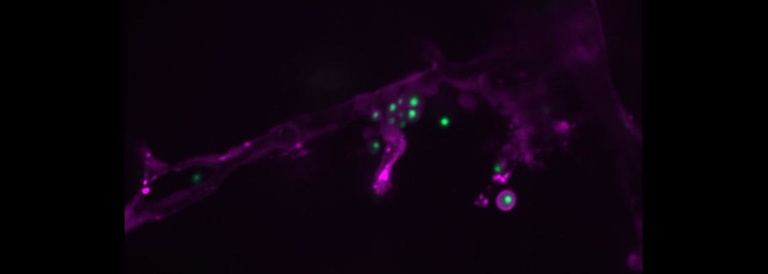
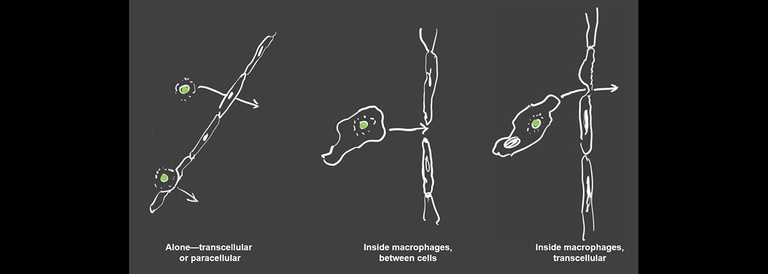
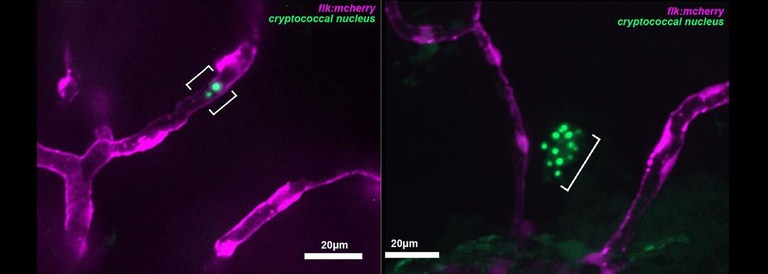
A key stage in the pathogenesis of Cryptococcus is crossing from the blood into the brain parenchyma to cause meningoencephalitis. The zebrafish larva provides an ideal model in which to observe and interrogate this process. A number of cryptococcal mutants have been found which are deficient in the process, but until now there has not been a unified way to investigate their specific roles and relationships all in the same fungal strain. We are using CRISPr technology to create a uniform library of these mutants which for direct study in the larva, which can then be used in combination to outline the pathways and chronology of this important event.
Granulomas and Granulomatous Inflammation
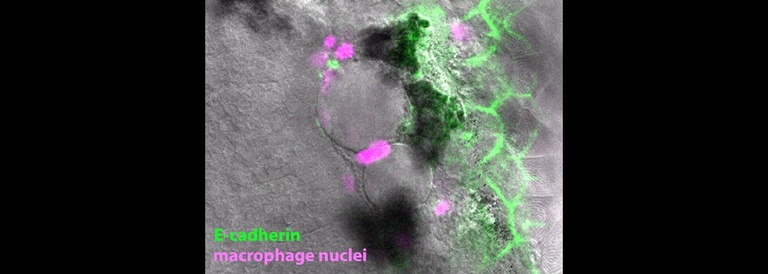
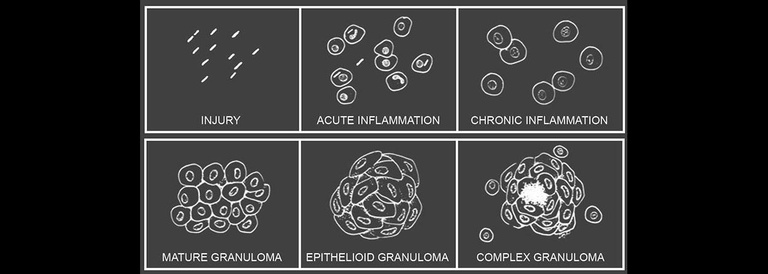
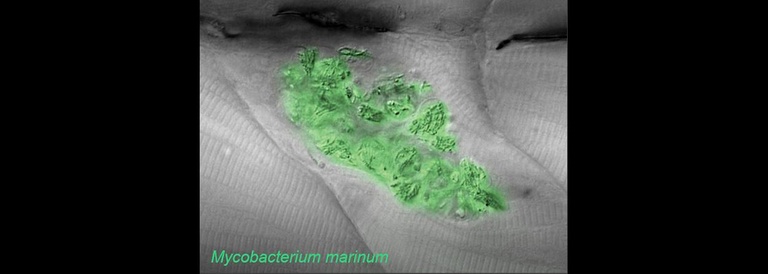
The word ‘granuloma’ refers to a pathological structure made up most essentially of macrophages and developing in the context of many forms of chronic inflammation. Granulomas form in a wide variety of human conditions from infections to cancers to autoinflammatory diseases—even in the context of thorns or other foreign bodies. A key goal of the lab is to define the earliest inflammatory and morphologic changes which are common to all forms of granulomatous inflammation. For this purpose we study host responses to several different stimuli, both infectious and inert. Granulomas formed in response to these varying triggers have widely divergent fates but begin with the same fundamental organization.
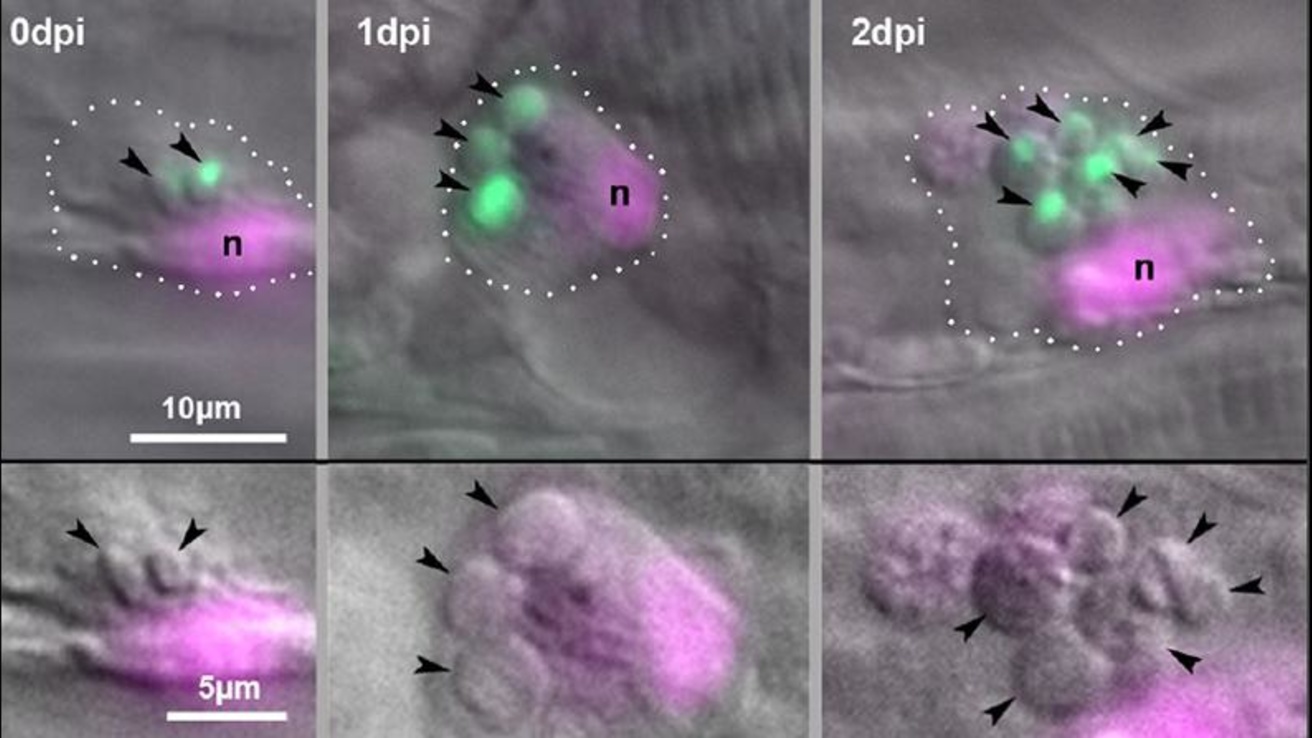
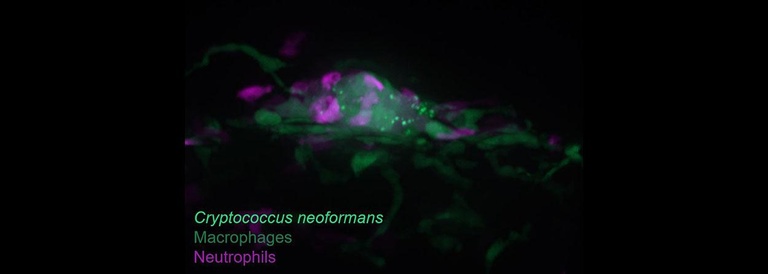
The Role of Germination in Cryptococcal Virulence and Persistence
The pathogenic Cryptococcus species are readily found in the environment and typically enter the body via inhalation. Clinical studies demonstrate that Cryptococcus neoformans can persist asymptomatically in humans for years, presumably inside granulomas. The yeast form is most common in the environment, but these species also form spores through both sexual and asexual processes. It remains uncertain whether it is the yeast or the spore form (or both) that acts as the propagule for human infection. Our studies in the zebrafish demonstrate that spore infection proceeds very differently from yeast infection, most notably in terms of granuloma formation and persistent, non-progressive infection. We are probing the processes of germination to learn more about how spores persist and induce granulomas. From Davis et al 2016.